Note: If you are looking for MeXpose v1 you can find it under the mexpose_v1 branch.
MeXpose 2.0 is a major update to our interactive workflow for comprehensive analysis of multiplexed imaging data, optimised for spatial metallomics. The new release streamlines the existing pipeline by replacing third-party software with custom Jupyter notebooks for image reconstruction, preprocessing, and data extraction. Additional new features such as limit-of-detection calculation, cytometry-style gating, and integrated spatial statistics enhance both exploratory and quantitative analyses within a single, intuitive environment.
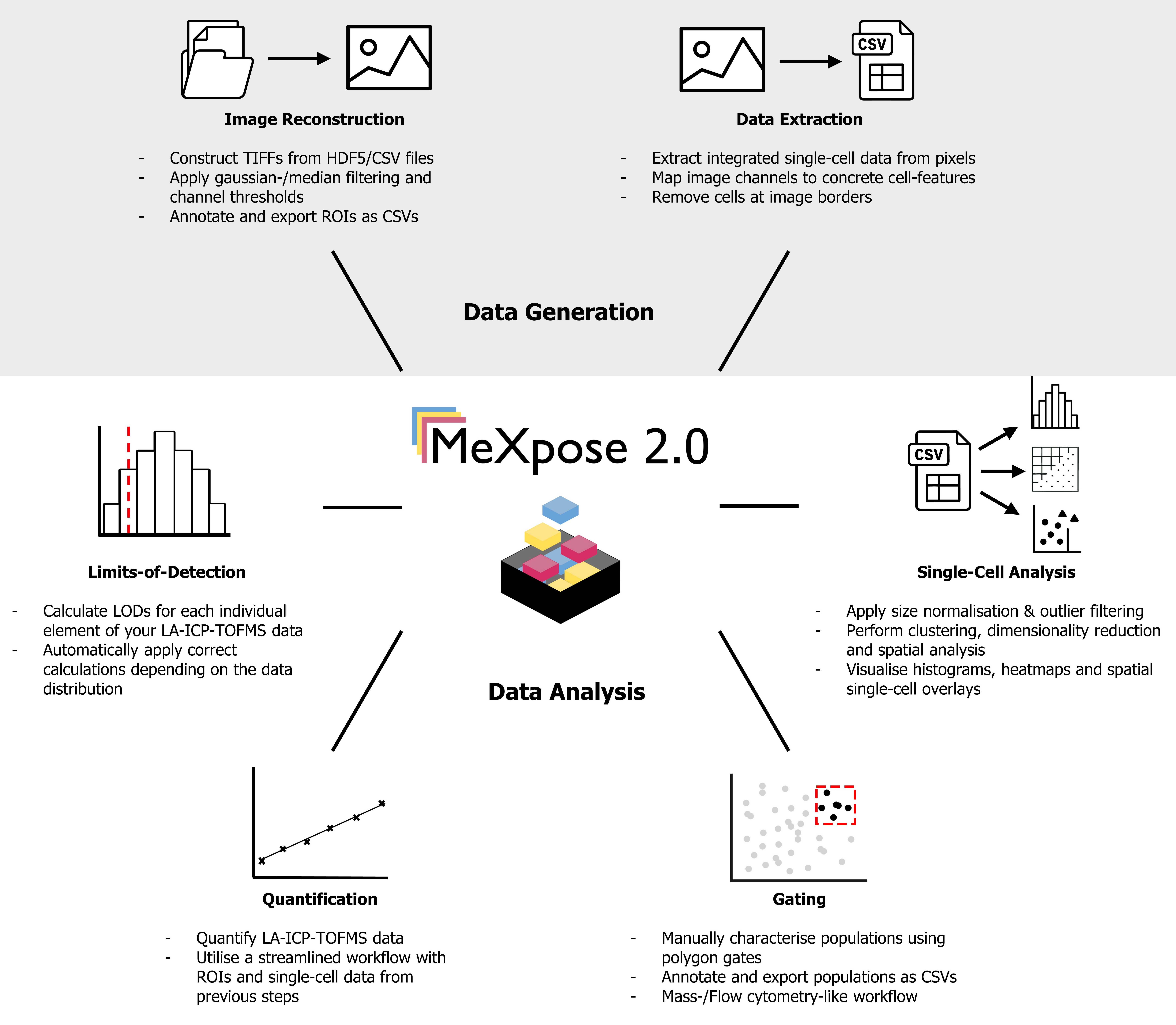
MeXpose 2.0 covers the following aspects of multiplexed image analysis by providing custom notebooks for each step:
- Image reconstruction (TOFWerk and Nu Instruments specific)
- Single-cell data extraction
- Single-cell analysis
- size normalisation & outlier filtering
- clustering, dimensionality reduction and spatial analysis
- histograms, heatmaps and spatial single-cell overlays
- Manual population gating
- Quantification
- Limit-of-detection calculations
All dependencies can be installed by creating a new mamba/conda environment using the provided mexpose_v2.yaml file:
# Create the environment from the yaml file
mamba env create -f mexpose_v2.yaml
# Activate the environment
mamba activate mexpose_v2Note: Replace mamba with conda when using Anaconda.
To handle large file uploads (>10 MB) in the notebooks, you'll need to create a custom JupyterLab configuration:
-
Create a JupyterLab config file:
jupyter lab --generate-config
-
Edit the configuration file (located at
~/.jupyter/jupyter_lab_config.py) and add:c.ServerApp.tornado_settings = {"websocket_max_message_size": 1024 * 1024 * 1024}
-
Create a JupyterLab config file:
jupyter lab --generate-config
-
Edit the configuration file (located at
%USERPROFILE%\.jupyter\jupyter_lab_config.py) and add:c.ServerApp.tornado_settings = {"websocket_max_message_size": 1024 * 1024 * 1024}
You can use the provided jupyter_lab_config.py file:
jupyter lab --config=jupyter_lab_config.pyThis increases the upload limit to 1 GB, allowing you to work with larger imaging datasets directly within the notebooks.
MeXpose 2.0 was tested on the following setups:
TOFWerk
| Software | Version | Manufacturer |
|---|---|---|
| TOFPilot | 2.15 | TOFWERK, Thun, Switzerland |
| Chromium | 3.2 | Teledyne Photon Machines, Bozeman, USA |
| ActiveView2 | 1.5.1.30 | Elemental Scientific Lasers (ESL), Bozeman, USA |
Using raw combined laser- and TOF data in HDF5 format
Nu Instruments
| Software | Version | Manufacturer |
|---|---|---|
| NuQuant | 1.2.8739.1 | Nu Instruments Ltd., Wrexham, UK |
| Chromium | 3.2 | Teledyne Photon Machines, Bozeman, USA |
| ActiveView2 | 1.5.1.30 | Elemental Scientific Lasers (ESL), Bozeman, USA |
Using preprocessed (NuQuant) combined laser- and TOF data in CSV format
Supported file types
- HDF5
- CSV
- TIFF, OME.TIFF, BIGTIFF, PNG, JPEG
Support for additional file formats is provided by the use of the imageio Python library.
MeXpose is focused around single-cell analysis. In order to extract single-cell data as well as for several visualisations a greyscale segmentation mask is required.
We recommend using (a recent version of) Cellpose as we find it performs very well and works reliably while additionally allowing annotation and training in a user-friendly GUI.
Visit the Cellpose github for installation instructions and their documentation for additional information.
You can also try Cellpose-SAM here:
Pachitariu, M., Rariden, M., & Stringer, C. (2025). Cellpose-SAM: superhuman generalization for cellular segmentation. bioRxiv.
Please cite the following papers when using either the original MeXpose or MeXpose 2.0 in your work:
MeXpose 2.0
To be updated
MeXpose 1.0
MeXpose - A modular imaging pipeline for the quantitative assessment of cellular metal bioaccumulation. Gabriel Braun and Martin Schaier, Paulina Werner, Sarah Theiner, Juergen Zanghellini, Lukas Wisgrill, Nanna Fyhrquist, Gunda Koellensperger. JACS Au 2024 4 (6), 2197-2210; DOI: 10.1021/jacsau.4c00154
@article{Braun_Schaier2024,
author = {Braun, Gabriel and Schaier, Martin and Werner, Paulina and Theiner, Sarah and Zanghellini, Jürgen and Wisgrill, Lukas and Fyhrquist, Nanna and Koellensperger, Gunda},
title = {MeXpose─A Modular Imaging Pipeline for the Quantitative Assessment of Cellular Metal Bioaccumulation},
journal = {JACS Au},
volume = {4},
number = {6},
pages = {2197-2210},
year = {2024},
doi = {10.1021/jacsau.4c00154},
URL = {https://doi.org/10.1021/jacsau.4c00154},
eprint = {https://doi.org/10.1021/jacsau.4c00154}
}The code included with MeXpose 2.0 is distributed under an MIT license.
The code included with MeXpose 1.0, namely all python scripts, jupyter notebooks and fiji macros, is distributed under an MIT license. The MeXpose container hosted on DockerHub is distributed under a GPLv3 license.