This is the unofficial code of the arxiv paper Protein Representation Learning by Geometric Structure Pretraining published by Zhang, Zuobai, et al.
It mainly consists of pretraining modules and GeomEtry-Aware Relational graph convolution network.
Please check dependency in yaml file and install with the following command:
conda env create --file gearnet.yaml
My codes mainly based on multi-GPU usages like 4 GPU in data parallel. Please uses 4 GPU if you are available.
Pretrained dataset can be downloaded in the following liks:
https://alphafold.ebi.ac.uk/download
protein dir contains raw pdb data, processed processed graph data and raw contains the text file that name of raw pdb files.
For the clarification, we used Swiss-Prot(PDB files) for pretraining model.
Thanks to Intrinsic-Extrinsic Convolution and Polling for Learning on 3D Protein Structures, we utilizes the dataset in follwing links :
- Fold classificiation
https://drive.google.com/uc?export=download&id=1chZAkaZlEBaOcjHQ3OUOdiKZqIn36qar
- Reaction classification
https://drive.google.com/uc?export=download&id=1udP6_90WYkwkvL1LwqIAzf9ibegBJ8rI
Note that my codes are not taking the data path as training argument. Please make sure that data path corretly set to saved data directory.
Download the model weight in following links: https://drive.google.com/drive/folders/1vozsHqYyBoGLhCI0GmqBipqYspd3U8t6?usp=sharing
-
Based V1 message function training
python multiview_contrast.py --num_worker 14 --save_path path where want to save --filename save file name --model_type v1-
resume from check points
python multiview_contrast.py --num_worker 14 --save_path where you want to save --filename save file name --model_type v1 --resume True --resume_path check point (/data/project/rw/codingtest/saved_info/temp2/multiview_v2_ckps_1.pt)
-
-
Based V2 message function training
python multiview_contrast.py --num_worker 14 --save_path where you want to save --filename save file name --model_type v2-
resume from ckps
python multiview_contrast.py --num_worker 14 --save_path where you want to save --filename save file name --model_type v2 --resume True --resume_path /data/project/rw/codingtest/saved_info/temp2/multiview_v2_ckps_2.pt
-
-
Based V1 message function training
python self_training_type1.py --num_worker 14 --save_path where you want to save --filename save file name --model_type v1-
resume from check points
python self_training_type1.py --num_worker 14 --save_path /data/project/rw/codingtest/saved_info/temp3/ --filename residue_type_pred_v1 --model_type v1 --resume True --resume_path check point (/data/project/rw/codingtest/saved_info/temp2/residue_pred_v1_ckps_1.pt)
-
-
Based V2 message function training
python self_training_type1.py --num_worker 14 --save_path where you want to save --filename save file name --model_type v2-
resume from check points
python self_training_type1.py --num_worker 14 --save_path /data/project/rw/codingtest/saved_info/temp3/ --filename residue_type_pred_v1 --model_type v2 --resume True --resume_path check point (/data/project/rw/codingtest/saved_info/temp2/residue_pred_v2_ckps_1.pt)
-
According to the paper, downstream task batch should 2 per GPU. I strongly recommend to use 4 GPU. Also, my codes automatically tries to load self-trained model weight at the initial step. Please set to the "multive_pretrained_epoch33.pt" in the correct saved path.
Notes that V2 model is not pretrained yet and I validate architectures in fold classifciation task with follwoing configuration.
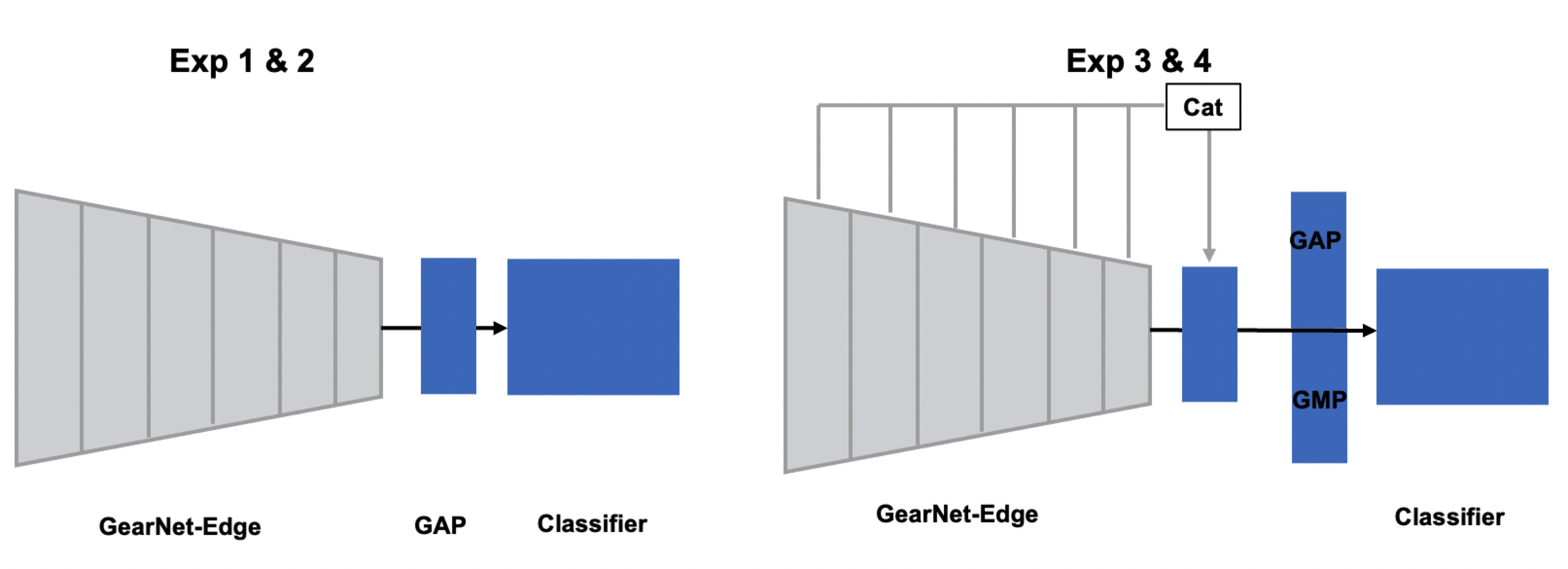
- Fold classification
-
train For V1 model with decoder 1
python fine_tune_fold.py --save_path /data/project/rw/codingtest/saved_info/temp3/fold_task/For V1 model with decoder 2
python fine_tune_fold.py --save_path /data/project/rw/codingtest/saved_info/temp3/fold_task/ --model_type v1 --decoder_pooling TrueFor V2 model with docoder 2
python fine_tune_fold.py --save_path /data/project/rw/codingtest/saved_info/temp3/fold_task/ --model_type v1 --decoder_pooling True -
test
For Experiment 1python fine_tune_fold.py --test True --resume_path ./modelweight/exp1.pt --model_type v1For Experiment 2
python fine_tune_fold.py --test True --resume_path ./modelweight/exp2.pt --model_type v1For Experiment 3
python fine_tune_fold.py --test True --resume_path ./modelweight/exp3.pt --model_type v1 --decoder_pooling TrueFor Experiment 4
python fine_tune_fold.py --test True --resume_path ./modelweight/exp3.pt --model_type v1 --decoder_pooling True
-
- Reaction classification
-
train & test This is same setting with above fine tune fold task.
python fine_tune_reaction.py --save_path /data/project/rw/codingtest/saved_info/temp4/fold_task/ --model_type v2 --decoder_pooling True
python fine_tune_fold.py --test True --resume_path ./modelweight/exp2.pt --model_type v1
-
I would appreciate if you found error. Please contact me via email address 구일kthan@gmail.com