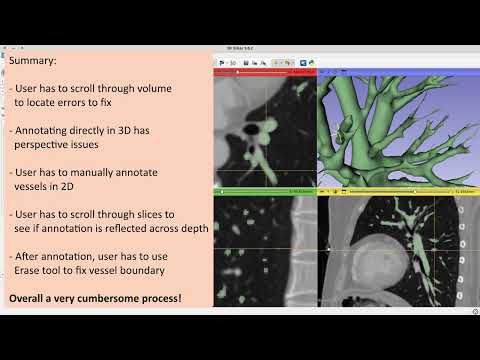
A 3D Slicer module for visualizing structure-wise uncertainty of vessels segmented by a deep learning model. The functionality involves adding, deleting, and viewing individual components based on their uncertainty values.
- 3D Slicer installed on your system
-
Clone the repository
git clone https://github.com/your-username/3d-slicer-uncertainty-visualizer.git
-
Enable Developer Mode
- Go to Slicer menu → Edit → Application Settings → Developer
- Check "Enable Developer Mode"
-
Load the Extension
- Navigate to Modules → Developer Tools → Extension Wizard
- Click 'Select Extension'
- Select the root folder of this GitHub repository
- Select New as the module to load
-
Access the Module
- Once the extension and module load successfully, go to Modules → Examples → New
Before clicking "Generate Uncertainty Map", ensure you have loaded these 3 files as Volumes:
-
Uncertainty File
- Rename this file following the pattern:
*uncertainty_file*before loading - Load as Volume (use the float volume, not the int version)
- Rename this file following the pattern:
-
Segmentation File
- Rename this file following the pattern:
*segmentation_file*before loading - Load as Volume
- Rename this file following the pattern:
-
Image File
- Load for visualization purposes
Important: Make sure no other files are loaded in Slicer that follow the above naming patterns.
- Click the "Generate Uncertainty Map" button to begin working
- The 'View' button for the 4th view (3D render) only works when the 4th view is in full screen mode
- Most of the code is located in
New.py